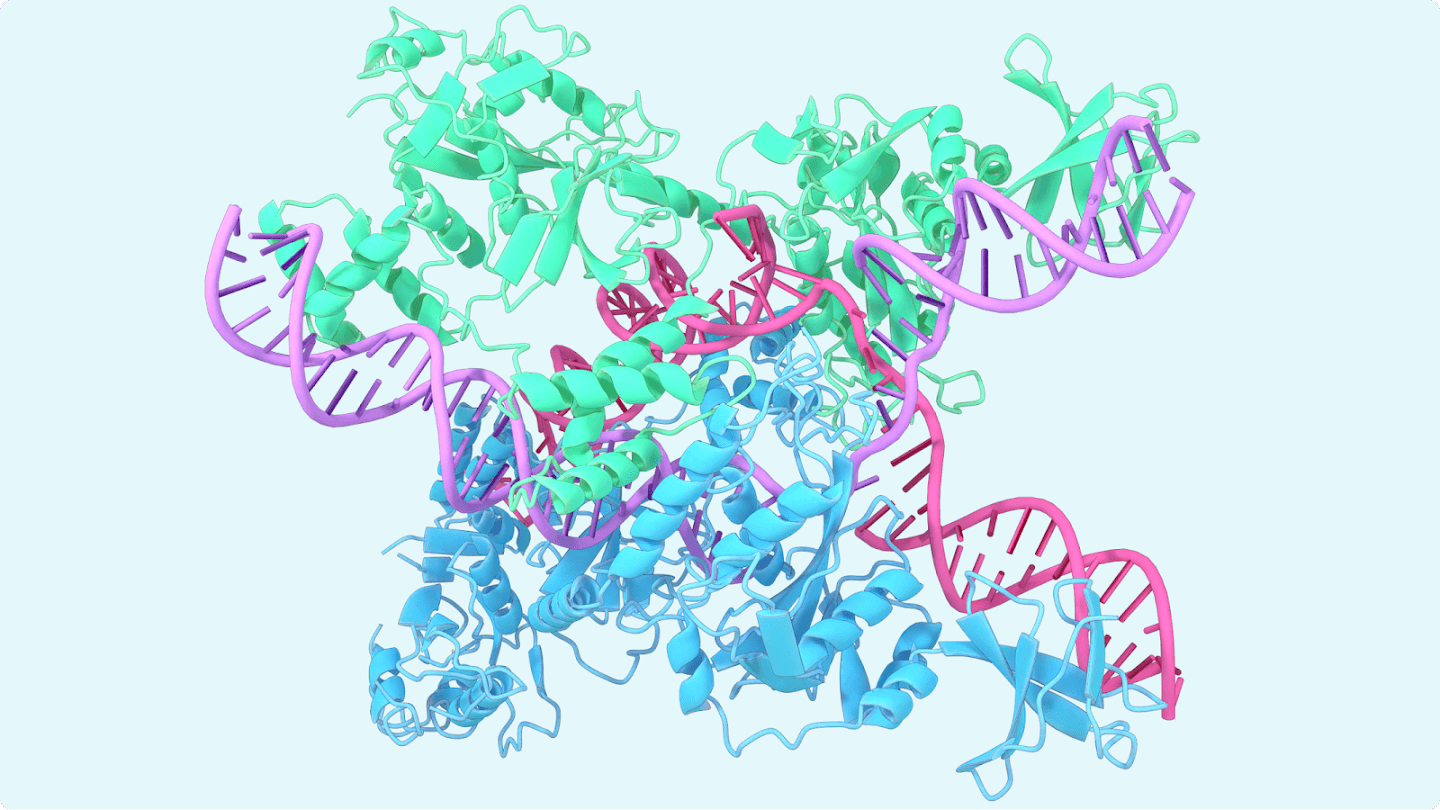
LONDON — Five years after AlphaFold stunned biologists by making high-accuracy protein structure prediction routine, Google DeepMind and its drug-discovery spinoff Isomorphic Labs are pushing the system into broader biology with AlphaFold 3, a model that can predict how proteins interact with DNA, RNA and many druglike molecules, Dec. 24, 2025.
The shift could speed early-stage research and drug design, but it has also reignited debate over how much of a scientific “public good” AlphaFold should be when the most capable versions are controlled through restricted releases.
AlphaFold 3 widens the lens beyond proteins
AlphaFold began as a solution to a decades-old bottleneck: turning a protein’s amino-acid sequence into a reliable 3D shape. AlphaFold 3 extends that promise to entire molecular complexes, including nucleic acids, small molecules, ions and modified residues, according to the peer-reviewed description in Nature. That matters because many of biology’s hardest problems — from gene regulation to drug binding — happen at the interfaces between molecules, not in a protein alone.
Developers have pitched AlphaFold 3 as a practical tool for drug discovery, saying it can help researchers reason about binding and interactions that are expensive to test at scale, as outlined in an Isomorphic Labs announcement. The work builds on AlphaFold’s growing role as infrastructure: the AlphaFold Protein Structure Database now offers open access to hundreds of millions of predicted protein structures.
From breakthrough to Nobel recognition
The Nobel committee credited AlphaFold’s impact when it awarded the 2024 Nobel Prize in chemistry to David Baker, Demis Hassabis and John Jumper, citing how the AI system helped crack the protein-structure prediction problem and opened new routes for protein design and discovery, according to the official Nobel Prize press release.
DeepMind has framed the five-year arc as a rapid migration from a competition-winning method to a widely used scientific tool, describing AlphaFold’s adoption and database milestones in a 2025 retrospective.
Why AlphaFold access is still contentious
The controversy is less about what AlphaFold 3 can do than who can scrutinize and reproduce it. When AlphaFold 3 was first published, researchers criticized the limited ability to run the model independently, a dispute that spilled into broader questions about journal standards and reproducibility, highlighted in Nature’s editorial explanation. DeepMind later moved toward wider availability for researchers, but access has remained more constrained than earlier releases, and some scientists argue that the shift slows verification and downstream innovation.
Continuity: the open era that set expectations
That tension is sharper because AlphaFold’s earlier chapters established a norm of openness. In 2021, DeepMind and EMBL-EBI launched a public database of predictions covering the human proteome and other organisms, described in an EMBL-EBI press release. A year later, a scientific overview in Nucleic Acids Research detailed the database’s expansion and programmatic access. And reporting in Wired captured how quickly the tool began reshaping everyday lab work.
Whether AlphaFold’s next five years are defined more by scientific reach or by governance and access may depend on how the field balances commercial drug-development ambitions with the openness that helped make AlphaFold a global standard in the first place.